[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user asfgit closed the pull request at: https://github.com/apache/spark/pull/22320 --- - To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user cloud-fan commented on a diff in the pull request: https://github.com/apache/spark/pull/22320#discussion_r215479502 --- Diff: sql/hive/src/main/scala/org/apache/spark/sql/hive/execution/CreateHiveTableAsSelectCommand.scala --- @@ -82,7 +83,7 @@ case class CreateHiveTableAsSelectCommand( query, overwrite = true, ifPartitionNotExists = false, - outputColumns = outputColumns).run(sparkSession, child) + outputColumnNames = outputColumnNames).run(sparkSession, child) --- End diff -- I feel it's better to specify parameters by name if the previous parameter is already specified by name, e.g. `ifPartitionNotExists = false` --- - To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user jaceklaskowski commented on a diff in the pull request: https://github.com/apache/spark/pull/22320#discussion_r215376132 --- Diff: sql/hive/src/main/scala/org/apache/spark/sql/hive/execution/CreateHiveTableAsSelectCommand.scala --- @@ -82,7 +83,7 @@ case class CreateHiveTableAsSelectCommand( query, overwrite = true, ifPartitionNotExists = false, - outputColumns = outputColumns).run(sparkSession, child) + outputColumnNames = outputColumnNames).run(sparkSession, child) --- End diff -- `outputColumnNames` themselves. Specyfing `outputColumnNames` as the name of the property to set using `outputColumnNames` does nothing but introduces a duplication. If you removed one `outputColumnNames` the comprehension should not be lowered whatsoever, shouldn't it? --- - To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user cloud-fan commented on a diff in the pull request: https://github.com/apache/spark/pull/22320#discussion_r215248202 --- Diff: sql/hive/src/main/scala/org/apache/spark/sql/hive/execution/CreateHiveTableAsSelectCommand.scala --- @@ -82,7 +83,7 @@ case class CreateHiveTableAsSelectCommand( query, overwrite = true, ifPartitionNotExists = false, - outputColumns = outputColumns).run(sparkSession, child) + outputColumnNames = outputColumnNames).run(sparkSession, child) --- End diff -- what's the duplication? --- - To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user cloud-fan commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r215247634
--- Diff:
sql/hive/src/test/scala/org/apache/spark/sql/hive/execution/HiveDDLSuite.scala
---
@@ -754,6 +754,54 @@ class HiveDDLSuite
}
}
+ test("Insert overwrite Hive table should output correct schema") {
+withSQLConf(CONVERT_METASTORE_PARQUET.key -> "false") {
+ withTable("tbl", "tbl2") {
+withView("view1") {
+ spark.sql("CREATE TABLE tbl(id long)")
+ spark.sql("INSERT OVERWRITE TABLE tbl VALUES 4")
--- End diff --
We can, but it's important to keep the code style consistent with the
existing code in the same file. In this test suite, seems SQL statements are
prefered.
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user cloud-fan commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r215246692
--- Diff:
sql/core/src/test/scala/org/apache/spark/sql/test/DataFrameReaderWriterSuite.scala
---
@@ -805,6 +805,80 @@ class DataFrameReaderWriterSuite extends QueryTest
with SharedSQLContext with Be
}
}
+ test("Insert overwrite table command should output correct schema:
basic") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).toDF("id")
--- End diff --
I think @gengliangwang meant case preserving, which is the behavior we are
testing against.
`spark.range(10).toDF("id")` is same as `spark.range(10)`, it's just
clearer to people who don't know `spark.range` outputs a single column named
"id".
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user jaceklaskowski commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r215215098
--- Diff:
sql/hive/src/test/scala/org/apache/spark/sql/hive/execution/HiveDDLSuite.scala
---
@@ -754,6 +754,54 @@ class HiveDDLSuite
}
}
+ test("Insert overwrite Hive table should output correct schema") {
+withSQLConf(CONVERT_METASTORE_PARQUET.key -> "false") {
+ withTable("tbl", "tbl2") {
+withView("view1") {
+ spark.sql("CREATE TABLE tbl(id long)")
+ spark.sql("INSERT OVERWRITE TABLE tbl VALUES 4")
--- End diff --
I might be missing something, but why does this test use SQL statements not
DataFrameWriter API, e.g.
`Seq(4).toDF("id").write.mode(SaveMode.Overwrite).saveAsTable("tbl")`?
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user jaceklaskowski commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r215213849
--- Diff:
sql/core/src/test/scala/org/apache/spark/sql/test/DataFrameReaderWriterSuite.scala
---
@@ -805,6 +805,80 @@ class DataFrameReaderWriterSuite extends QueryTest
with SharedSQLContext with Be
}
}
+ test("Insert overwrite table command should output correct schema:
basic") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).toDF("id")
--- End diff --
"case sensitive"? How is so since Spark SQL is case-insensitive by default?
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user jaceklaskowski commented on a diff in the pull request: https://github.com/apache/spark/pull/22320#discussion_r215214259 --- Diff: sql/hive/src/main/scala/org/apache/spark/sql/hive/execution/CreateHiveTableAsSelectCommand.scala --- @@ -82,7 +83,7 @@ case class CreateHiveTableAsSelectCommand( query, overwrite = true, ifPartitionNotExists = false, - outputColumns = outputColumns).run(sparkSession, child) + outputColumnNames = outputColumnNames).run(sparkSession, child) --- End diff -- Why is this duplication needed here? --- - To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user gengliangwang commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r215128076
--- Diff:
sql/core/src/main/scala/org/apache/spark/sql/execution/datasources/InsertIntoHadoopFsRelationCommand.scala
---
@@ -56,7 +56,7 @@ case class InsertIntoHadoopFsRelationCommand(
mode: SaveMode,
catalogTable: Option[CatalogTable],
fileIndex: Option[FileIndex],
-outputColumns: Seq[Attribute])
+outputColumnNames: Seq[String])
extends DataWritingCommand {
import
org.apache.spark.sql.catalyst.catalog.ExternalCatalogUtils.escapePathName
--- End diff --
Oh, then we can use this method instead.
```
def checkColumnNameDuplication(
columnNames: Seq[String], colType: String, caseSensitiveAnalysis:
Boolean): Unit
```
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user wangyum commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r215106921
--- Diff:
sql/core/src/main/scala/org/apache/spark/sql/execution/datasources/InsertIntoHadoopFsRelationCommand.scala
---
@@ -56,7 +56,7 @@ case class InsertIntoHadoopFsRelationCommand(
mode: SaveMode,
catalogTable: Option[CatalogTable],
fileIndex: Option[FileIndex],
-outputColumns: Seq[Attribute])
+outputColumnNames: Seq[String])
extends DataWritingCommand {
import
org.apache.spark.sql.catalyst.catalog.ExternalCatalogUtils.escapePathName
--- End diff --
Line 66: `query.schema` should be
`DataWritingCommand.logicalPlanSchemaWithNames(query, outputColumnNames)`.
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user gengliangwang commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214828936
--- Diff:
sql/hive/src/main/scala/org/apache/spark/sql/hive/execution/CreateHiveTableAsSelectCommand.scala
---
@@ -63,13 +63,14 @@ case class CreateHiveTableAsSelectCommand(
query,
overwrite = false,
ifPartitionNotExists = false,
-outputColumns = outputColumns).run(sparkSession, child)
+outputColumnNames = outputColumnNames).run(sparkSession, child)
} else {
// TODO ideally, we should get the output data ready first and then
// add the relation into catalog, just in case of failure occurs
while data
// processing.
assert(tableDesc.schema.isEmpty)
- catalog.createTable(tableDesc.copy(schema = query.schema),
ignoreIfExists = false)
+ val schema = DataWritingCommand.logicalPlanSchemaWithNames(query,
outputColumnNames)
+ catalog.createTable(tableDesc.copy(schema = schema), ignoreIfExists
= false)
--- End diff --
The schema naming need to be consistent with `outputColumnNames` here.
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user gengliangwang commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214828496
--- Diff:
sql/hive/src/test/scala/org/apache/spark/sql/hive/execution/HiveDDLSuite.scala
---
@@ -754,6 +754,47 @@ class HiveDDLSuite
}
}
+ test("Insert overwrite Hive table should output correct schema") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+spark.sql("CREATE TABLE tbl(id long)")
+spark.sql("INSERT OVERWRITE TABLE tbl SELECT 4")
+spark.sql("CREATE VIEW view1 AS SELECT id FROM tbl")
+spark.sql("CREATE TABLE tbl2(ID long)")
+spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT ID FROM view1")
+checkAnswer(spark.table("tbl2"), Seq(Row(4)))
--- End diff --
Good point. I found that `CreateHiveTableAsSelectCommand` output wrong
schema after adding a new test case.
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user wangyum commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214786494
--- Diff:
sql/hive/src/test/scala/org/apache/spark/sql/hive/execution/HiveDDLSuite.scala
---
@@ -754,6 +754,47 @@ class HiveDDLSuite
}
}
+ test("Insert overwrite Hive table should output correct schema") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+spark.sql("CREATE TABLE tbl(id long)")
+spark.sql("INSERT OVERWRITE TABLE tbl SELECT 4")
+spark.sql("CREATE VIEW view1 AS SELECT id FROM tbl")
+spark.sql("CREATE TABLE tbl2(ID long)")
+spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT ID FROM view1")
+checkAnswer(spark.table("tbl2"), Seq(Row(4)))
--- End diff --
Add schema assert please. We can read data since
[SPARK-25132](https://issues.apache.org/jira/browse/SPARK-25132).
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user gengliangwang commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214778690
--- Diff:
sql/core/src/test/scala/org/apache/spark/sql/test/DataFrameReaderWriterSuite.scala
---
@@ -805,6 +805,80 @@ class DataFrameReaderWriterSuite extends QueryTest
with SharedSQLContext with Be
}
}
+ test("Insert overwrite table command should output correct schema:
basic") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).toDF("id")
+df.write.format("parquet").saveAsTable("tbl")
+spark.sql("CREATE VIEW view1 AS SELECT id FROM tbl")
+spark.sql("CREATE TABLE tbl2(ID long) USING parquet")
+spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT ID FROM view1")
+val identifier = TableIdentifier("tbl2", Some("default"))
+val location =
spark.sessionState.catalog.getTableMetadata(identifier).location.toString
+val expectedSchema = StructType(Seq(StructField("ID", LongType,
true)))
+assert(spark.read.parquet(location).schema == expectedSchema)
+checkAnswer(spark.table("tbl2"), df)
+ }
+}
+ }
+
+ test("Insert overwrite table command should output correct schema:
complex") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).map(x => (x, x.toInt,
x.toInt)).toDF("col1", "col2", "col3")
+df.write.format("parquet").saveAsTable("tbl")
+spark.sql("CREATE VIEW view1 AS SELECT * FROM tbl")
+spark.sql("CREATE TABLE tbl2(COL1 long, COL2 int, COL3 int) USING
parquet PARTITIONED " +
+ "BY (COL2) CLUSTERED BY (COL3) INTO 3 BUCKETS")
+spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT COL1, COL2, COL3
FROM view1")
+val identifier = TableIdentifier("tbl2", Some("default"))
+val location =
spark.sessionState.catalog.getTableMetadata(identifier).location.toString
+val expectedSchema = StructType(Seq(
+ StructField("COL1", LongType, true),
--- End diff --
Keep it should be OK.
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user gengliangwang commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214778523
--- Diff:
sql/core/src/test/scala/org/apache/spark/sql/test/DataFrameReaderWriterSuite.scala
---
@@ -805,6 +805,80 @@ class DataFrameReaderWriterSuite extends QueryTest
with SharedSQLContext with Be
}
}
+ test("Insert overwrite table command should output correct schema:
basic") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).toDF("id")
--- End diff --
This is trivial...As the column name `id` is case sensitive and used below,
I would like to show it explicitly.
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user maropu commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214761843
--- Diff:
sql/hive/src/main/scala/org/apache/spark/sql/hive/execution/InsertIntoHiveTable.scala
---
@@ -69,7 +69,7 @@ case class InsertIntoHiveTable(
query: LogicalPlan,
overwrite: Boolean,
ifPartitionNotExists: Boolean,
-outputColumns: Seq[Attribute]) extends SaveAsHiveFile {
+outputColumnNames: Seq[String]) extends SaveAsHiveFile {
--- End diff --
thanks!
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user jaceklaskowski commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214751309
--- Diff:
sql/core/src/test/scala/org/apache/spark/sql/test/DataFrameReaderWriterSuite.scala
---
@@ -805,6 +805,80 @@ class DataFrameReaderWriterSuite extends QueryTest
with SharedSQLContext with Be
}
}
+ test("Insert overwrite table command should output correct schema:
basic") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).toDF("id")
+df.write.format("parquet").saveAsTable("tbl")
+spark.sql("CREATE VIEW view1 AS SELECT id FROM tbl")
+spark.sql("CREATE TABLE tbl2(ID long) USING parquet")
+spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT ID FROM view1")
+val identifier = TableIdentifier("tbl2", Some("default"))
+val location =
spark.sessionState.catalog.getTableMetadata(identifier).location.toString
+val expectedSchema = StructType(Seq(StructField("ID", LongType,
true)))
+assert(spark.read.parquet(location).schema == expectedSchema)
+checkAnswer(spark.table("tbl2"), df)
+ }
+}
+ }
+
+ test("Insert overwrite table command should output correct schema:
complex") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).map(x => (x, x.toInt,
x.toInt)).toDF("col1", "col2", "col3")
+df.write.format("parquet").saveAsTable("tbl")
+spark.sql("CREATE VIEW view1 AS SELECT * FROM tbl")
+spark.sql("CREATE TABLE tbl2(COL1 long, COL2 int, COL3 int) USING
parquet PARTITIONED " +
+ "BY (COL2) CLUSTERED BY (COL3) INTO 3 BUCKETS")
+spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT COL1, COL2, COL3
FROM view1")
+val identifier = TableIdentifier("tbl2", Some("default"))
+val location =
spark.sessionState.catalog.getTableMetadata(identifier).location.toString
+val expectedSchema = StructType(Seq(
+ StructField("COL1", LongType, true),
+ StructField("COL3", IntegerType, true),
--- End diff --
You could use a little magic here: `$"COL1".int`
```
scala> $"COL1".int
res1: org.apache.spark.sql.types.StructField =
StructField(COL1,IntegerType,true)
```
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user jaceklaskowski commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214750815
--- Diff:
sql/core/src/test/scala/org/apache/spark/sql/test/DataFrameReaderWriterSuite.scala
---
@@ -805,6 +805,80 @@ class DataFrameReaderWriterSuite extends QueryTest
with SharedSQLContext with Be
}
}
+ test("Insert overwrite table command should output correct schema:
basic") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).toDF("id")
--- End diff --
Why is `toDF("id")` required? Why not `spark.range(10)` alone?
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user jaceklaskowski commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214751930
--- Diff:
sql/hive/src/test/scala/org/apache/spark/sql/hive/execution/HiveDDLSuite.scala
---
@@ -754,6 +754,47 @@ class HiveDDLSuite
}
}
+ test("Insert overwrite Hive table should output correct schema") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+spark.sql("CREATE TABLE tbl(id long)")
+spark.sql("INSERT OVERWRITE TABLE tbl SELECT 4")
+spark.sql("CREATE VIEW view1 AS SELECT id FROM tbl")
+spark.sql("CREATE TABLE tbl2(ID long)")
+spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT ID FROM view1")
+checkAnswer(spark.table("tbl2"), Seq(Row(4)))
+ }
+}
+ }
+
+ test("Insert into Hive directory should output correct schema") {
+withTable("tbl") {
+ withView("view1") {
+withTempPath { path =>
+ spark.sql("CREATE TABLE tbl(id long)")
+ spark.sql("INSERT OVERWRITE TABLE tbl SELECT 4")
--- End diff --
`s/SELECT/VALUES` as it could be a bit more Spark-idiomatic?
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user jaceklaskowski commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214751219
--- Diff:
sql/core/src/test/scala/org/apache/spark/sql/test/DataFrameReaderWriterSuite.scala
---
@@ -805,6 +805,80 @@ class DataFrameReaderWriterSuite extends QueryTest
with SharedSQLContext with Be
}
}
+ test("Insert overwrite table command should output correct schema:
basic") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).toDF("id")
+df.write.format("parquet").saveAsTable("tbl")
+spark.sql("CREATE VIEW view1 AS SELECT id FROM tbl")
+spark.sql("CREATE TABLE tbl2(ID long) USING parquet")
+spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT ID FROM view1")
+val identifier = TableIdentifier("tbl2", Some("default"))
+val location =
spark.sessionState.catalog.getTableMetadata(identifier).location.toString
+val expectedSchema = StructType(Seq(StructField("ID", LongType,
true)))
+assert(spark.read.parquet(location).schema == expectedSchema)
+checkAnswer(spark.table("tbl2"), df)
+ }
+}
+ }
+
+ test("Insert overwrite table command should output correct schema:
complex") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).map(x => (x, x.toInt,
x.toInt)).toDF("col1", "col2", "col3")
+df.write.format("parquet").saveAsTable("tbl")
+spark.sql("CREATE VIEW view1 AS SELECT * FROM tbl")
+spark.sql("CREATE TABLE tbl2(COL1 long, COL2 int, COL3 int) USING
parquet PARTITIONED " +
+ "BY (COL2) CLUSTERED BY (COL3) INTO 3 BUCKETS")
+spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT COL1, COL2, COL3
FROM view1")
+val identifier = TableIdentifier("tbl2", Some("default"))
+val location =
spark.sessionState.catalog.getTableMetadata(identifier).location.toString
+val expectedSchema = StructType(Seq(
+ StructField("COL1", LongType, true),
--- End diff --
`nullable` is `true` by default.
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user jaceklaskowski commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214751023
--- Diff:
sql/core/src/test/scala/org/apache/spark/sql/test/DataFrameReaderWriterSuite.scala
---
@@ -805,6 +805,80 @@ class DataFrameReaderWriterSuite extends QueryTest
with SharedSQLContext with Be
}
}
+ test("Insert overwrite table command should output correct schema:
basic") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).toDF("id")
+df.write.format("parquet").saveAsTable("tbl")
+spark.sql("CREATE VIEW view1 AS SELECT id FROM tbl")
+spark.sql("CREATE TABLE tbl2(ID long) USING parquet")
+spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT ID FROM view1")
+val identifier = TableIdentifier("tbl2", Some("default"))
--- End diff --
`default` is the default database name, isn't it? I'd remove it from the
test or use `spark.catalog.currentDatabase`.
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user jaceklaskowski commented on a diff in the pull request: https://github.com/apache/spark/pull/22320#discussion_r214751748 --- Diff: sql/hive/src/main/scala/org/apache/spark/sql/hive/execution/CreateHiveTableAsSelectCommand.scala --- @@ -63,7 +63,7 @@ case class CreateHiveTableAsSelectCommand( query, overwrite = false, ifPartitionNotExists = false, -outputColumns = outputColumns).run(sparkSession, child) +outputColumnNames = outputColumnNames).run(sparkSession, child) --- End diff -- Can you remove one `outputColumnNames`? --- - To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user jaceklaskowski commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214751169
--- Diff:
sql/core/src/test/scala/org/apache/spark/sql/test/DataFrameReaderWriterSuite.scala
---
@@ -805,6 +805,80 @@ class DataFrameReaderWriterSuite extends QueryTest
with SharedSQLContext with Be
}
}
+ test("Insert overwrite table command should output correct schema:
basic") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).toDF("id")
+df.write.format("parquet").saveAsTable("tbl")
+spark.sql("CREATE VIEW view1 AS SELECT id FROM tbl")
+spark.sql("CREATE TABLE tbl2(ID long) USING parquet")
+spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT ID FROM view1")
+val identifier = TableIdentifier("tbl2", Some("default"))
+val location =
spark.sessionState.catalog.getTableMetadata(identifier).location.toString
+val expectedSchema = StructType(Seq(StructField("ID", LongType,
true)))
+assert(spark.read.parquet(location).schema == expectedSchema)
+checkAnswer(spark.table("tbl2"), df)
+ }
+}
+ }
+
+ test("Insert overwrite table command should output correct schema:
complex") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).map(x => (x, x.toInt,
x.toInt)).toDF("col1", "col2", "col3")
+df.write.format("parquet").saveAsTable("tbl")
+spark.sql("CREATE VIEW view1 AS SELECT * FROM tbl")
+spark.sql("CREATE TABLE tbl2(COL1 long, COL2 int, COL3 int) USING
parquet PARTITIONED " +
+ "BY (COL2) CLUSTERED BY (COL3) INTO 3 BUCKETS")
+spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT COL1, COL2, COL3
FROM view1")
+val identifier = TableIdentifier("tbl2", Some("default"))
--- End diff --
Same as above.
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user gengliangwang commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214735437
--- Diff:
sql/hive/src/test/scala/org/apache/spark/sql/hive/execution/HiveDDLSuite.scala
---
@@ -754,6 +754,47 @@ class HiveDDLSuite
}
}
+ test("Insert overwrite Hive table should output correct schema") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+spark.sql("CREATE TABLE tbl(id long)")
--- End diff --
I am not familiar with Hive. But as I look at the debug message of this
logical plan, the top level is `InsertIntoHiveTable `default`.`tbl2`,
org.apache.hadoop.hive.serde2.lazy.LazySimpleSerDe, true, false, [ID]`. It
should not be related to this configuration, right?
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user cloud-fan commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214722461
--- Diff:
sql/hive/src/test/scala/org/apache/spark/sql/hive/execution/HiveDDLSuite.scala
---
@@ -754,6 +754,47 @@ class HiveDDLSuite
}
}
+ test("Insert overwrite Hive table should output correct schema") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+spark.sql("CREATE TABLE tbl(id long)")
--- End diff --
please run this test within
`withSQLConf(HiveUtils.CONVERT_METASTORE_PARQUET -> false)`
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user cloud-fan commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214722030
--- Diff:
sql/core/src/test/scala/org/apache/spark/sql/test/DataFrameReaderWriterSuite.scala
---
@@ -805,6 +805,81 @@ class DataFrameReaderWriterSuite extends QueryTest
with SharedSQLContext with Be
}
}
+ test("Insert overwrite table command should output correct schema:
basic") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).toDF("id")
+df.write.format("parquet").saveAsTable("tbl")
+spark.sql("CREATE VIEW view1 AS SELECT id FROM tbl")
+spark.sql("CREATE TABLE tbl2(ID long) USING parquet")
+spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT ID FROM view1")
+val identifier = TableIdentifier("tbl2", Some("default"))
+val location =
spark.sessionState.catalog.getTableMetadata(identifier).location.toString
+val expectedSchema = StructType(Seq(StructField("ID", LongType,
true)))
+assert(spark.read.parquet(location).schema == expectedSchema)
+checkAnswer(spark.table("tbl2"), df)
+ }
+}
+ }
+
+ test("Insert overwrite table command should output correct schema:
complex") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).map(x => (x, x.toInt,
x.toInt)).toDF("col1", "col2", "col3")
+df.write.format("parquet").saveAsTable("tbl")
+spark.sql("CREATE VIEW view1 AS SELECT * FROM tbl")
+spark.sql("CREATE TABLE tbl2(COL1 long, COL2 int, COL3 int) USING
parquet PARTITIONED " +
+ "BY (COL2) CLUSTERED BY (COL3) INTO 3 BUCKETS")
+spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT COL1, COL2, COL3 " +
+ "FROM view1 CLUSTER BY COL3")
--- End diff --
is it legal to put `CLUSTER BY` in the INSERT statement?
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user cloud-fan commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214721624
--- Diff: sql/core/src/test/scala/org/apache/spark/sql/SQLQuerySuite.scala
---
@@ -24,6 +24,7 @@ import java.util.concurrent.atomic.AtomicBoolean
import org.apache.spark.{AccumulatorSuite, SparkException}
import org.apache.spark.scheduler.{SparkListener, SparkListenerJobStart}
+import org.apache.spark.sql.catalyst.TableIdentifier
--- End diff --
unnecessary change
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user gengliangwang commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214697039
--- Diff:
sql/core/src/main/scala/org/apache/spark/sql/execution/command/DataWritingCommand.scala
---
@@ -53,3 +57,21 @@ trait DataWritingCommand extends Command {
def run(sparkSession: SparkSession, child: SparkPlan): Seq[Row]
}
+
+object DataWritingCommand {
+ /**
+ * Returns output attributes with provided names.
+ * The length of provided names should be the same of the length of
[[LogicalPlan.output]].
+ */
+ def logicalPlanOutputWithNames(
+ query: LogicalPlan,
+ names: Seq[String]): Seq[Attribute] = {
+// Save the output attributes to a variable to avoid duplicated
function calls.
+val outputAttributes = query.output
--- End diff --
I think both are OK. The current way makes it easier to call this Util
function, while the ways you suggests makes the argument carrying minimal
information.
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user gengliangwang commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214694881
--- Diff:
sql/hive/src/main/scala/org/apache/spark/sql/hive/execution/InsertIntoHiveTable.scala
---
@@ -69,7 +69,7 @@ case class InsertIntoHiveTable(
query: LogicalPlan,
overwrite: Boolean,
ifPartitionNotExists: Boolean,
-outputColumns: Seq[Attribute]) extends SaveAsHiveFile {
+outputColumnNames: Seq[String]) extends SaveAsHiveFile {
--- End diff --
No problem ð
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user maropu commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214671722
--- Diff:
sql/hive/src/main/scala/org/apache/spark/sql/hive/execution/InsertIntoHiveTable.scala
---
@@ -69,7 +69,7 @@ case class InsertIntoHiveTable(
query: LogicalPlan,
overwrite: Boolean,
ifPartitionNotExists: Boolean,
-outputColumns: Seq[Attribute]) extends SaveAsHiveFile {
+outputColumnNames: Seq[String]) extends SaveAsHiveFile {
--- End diff --
For better test coverage, can you add tests for hive tables?
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user maropu commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214671466
--- Diff: sql/core/src/test/scala/org/apache/spark/sql/SQLQuerySuite.scala
---
@@ -2853,6 +2854,81 @@ class SQLQuerySuite extends QueryTest with
SharedSQLContext {
}
}
+ test("Insert overwrite table command should output correct schema:
basic") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).toDF("id")
+df.write.format("parquet").saveAsTable("tbl")
+spark.sql("CREATE VIEW view1 AS SELECT id FROM tbl")
+spark.sql("CREATE TABLE tbl2(ID long) USING parquet")
+spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT ID FROM view1")
+val identifier = TableIdentifier("tbl2", Some("default"))
+val location =
spark.sessionState.catalog.getTableMetadata(identifier).location.toString
+val expectedSchema = StructType(Seq(StructField("ID", LongType,
true)))
+assert(spark.read.parquet(location).schema == expectedSchema)
+checkAnswer(spark.table("tbl2"), df)
+ }
+}
+ }
+
+ test("Insert overwrite table command should output correct schema:
complex") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).map(x => (x, x.toInt,
x.toInt)).toDF("col1", "col2", "col3")
+df.write.format("parquet").saveAsTable("tbl")
+spark.sql("CREATE VIEW view1 AS SELECT * FROM tbl")
+spark.sql("CREATE TABLE tbl2(COL1 long, COL2 int, COL3 int) USING
parquet PARTITIONED " +
+ "BY (COL2) CLUSTERED BY (COL3) INTO 3 BUCKETS")
+spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT COL1, COL2, COL3 " +
+ "FROM view1 CLUSTER BY COL3")
+val identifier = TableIdentifier("tbl2", Some("default"))
+val location =
spark.sessionState.catalog.getTableMetadata(identifier).location.toString
+val expectedSchema = StructType(Seq(
+ StructField("COL1", LongType, true),
+ StructField("COL3", IntegerType, true),
+ StructField("COL2", IntegerType, true)))
+assert(spark.read.parquet(location).schema == expectedSchema)
+checkAnswer(spark.table("tbl2"), df)
+ }
+}
+ }
+
+ test("Create table as select command should output correct schema:
basic") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).toDF("id")
+df.write.format("parquet").saveAsTable("tbl")
+spark.sql("CREATE VIEW view1 AS SELECT id FROM tbl")
+spark.sql("CREATE TABLE tbl2 USING parquet AS SELECT ID FROM
view1")
+val identifier = TableIdentifier("tbl2", Some("default"))
+val location =
spark.sessionState.catalog.getTableMetadata(identifier).location.toString
+val expectedSchema = StructType(Seq(StructField("ID", LongType,
true)))
+assert(spark.read.parquet(location).schema == expectedSchema)
+checkAnswer(spark.table("tbl2"), df)
+ }
+}
+ }
+
+ test("Create table as select command should output correct schema:
complex") {
+withTable("tbl", "tbl2") {
+ withView("view1") {
+val df = spark.range(10).map(x => (x, x.toInt,
x.toInt)).toDF("col1", "col2", "col3")
+df.write.format("parquet").saveAsTable("tbl")
+spark.sql("CREATE VIEW view1 AS SELECT * FROM tbl")
+spark.sql("CREATE TABLE tbl2 USING parquet PARTITIONED BY (COL2) "
+
+ "CLUSTERED BY (COL3) INTO 3 BUCKETS AS SELECT COL1, COL2, COL3
FROM view1")
+val identifier = TableIdentifier("tbl2", Some("default"))
+val location =
spark.sessionState.catalog.getTableMetadata(identifier).location.toString
+val expectedSchema = StructType(Seq(
+ StructField("COL1", LongType, true),
+ StructField("COL3", IntegerType, true),
+ StructField("COL2", IntegerType, true)))
+assert(spark.read.parquet(location).schema == expectedSchema)
+checkAnswer(spark.table("tbl2"), df)
+ }
+}
+ }
+
--- End diff --
better to move these tests into `DataFrameReaderWriterSuite`?
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user maropu commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214658233
--- Diff:
sql/core/src/main/scala/org/apache/spark/sql/execution/command/DataWritingCommand.scala
---
@@ -53,3 +57,21 @@ trait DataWritingCommand extends Command {
def run(sparkSession: SparkSession, child: SparkPlan): Seq[Row]
}
+
+object DataWritingCommand {
+ /**
+ * Returns output attributes with provided names.
+ * The length of provided names should be the same of the length of
[[LogicalPlan.output]].
+ */
+ def logicalPlanOutputWithNames(
+ query: LogicalPlan,
+ names: Seq[String]): Seq[Attribute] = {
+// Save the output attributes to a variable to avoid duplicated
function calls.
+val outputAttributes = query.output
--- End diff --
`query: LogicalPlan` -> `outputAttributes: Seq[Attribute]` in the function
argument, then drop the line above?
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user gengliangwang commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214655750
--- Diff:
sql/catalyst/src/main/scala/org/apache/spark/sql/catalyst/plans/QueryPlan.scala
---
@@ -38,6 +38,20 @@ abstract class QueryPlan[PlanType <:
QueryPlan[PlanType]] extends TreeNode[PlanT
*/
def outputSet: AttributeSet = AttributeSet(output)
+ /**
+ * Returns output attributes with provided names.
+ * The length of provided names should be the same of the length of
[[output]].
+ */
+ def outputWithNames(names: Seq[String]): Seq[Attribute] = {
+// Save the output attributes to a variable to avoid duplicated
function calls.
+val outputAttributes = output
+assert(outputAttributes.length == names.length,
+ "The length of provided names doesn't match the length of output
attributes.")
+outputAttributes.zipWithIndex.map { case (element, index) =>
+ element.withName(names(index))
+}
+ }
+
--- End diff --
@maropu Thanks! I have create object `DataWritingCommand` for this.
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user maropu commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214655488
--- Diff:
sql/catalyst/src/main/scala/org/apache/spark/sql/catalyst/plans/QueryPlan.scala
---
@@ -38,6 +38,20 @@ abstract class QueryPlan[PlanType <:
QueryPlan[PlanType]] extends TreeNode[PlanT
*/
def outputSet: AttributeSet = AttributeSet(output)
+ /**
+ * Returns output attributes with provided names.
+ * The length of provided names should be the same of the length of
[[output]].
+ */
+ def outputWithNames(names: Seq[String]): Seq[Attribute] = {
+// Save the output attributes to a variable to avoid duplicated
function calls.
+val outputAttributes = output
+assert(outputAttributes.length == names.length,
+ "The length of provided names doesn't match the length of output
attributes.")
+outputAttributes.zipWithIndex.map { case (element, index) =>
+ element.withName(names(index))
--- End diff --
`outputAttributes.zip(names).map { case (attr, outputName) =>
attr.withName(outputName) }`?
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user maropu commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214653309
--- Diff:
sql/catalyst/src/main/scala/org/apache/spark/sql/catalyst/plans/QueryPlan.scala
---
@@ -38,6 +38,20 @@ abstract class QueryPlan[PlanType <:
QueryPlan[PlanType]] extends TreeNode[PlanT
*/
def outputSet: AttributeSet = AttributeSet(output)
+ /**
+ * Returns output attributes with provided names.
+ * The length of provided names should be the same of the length of
[[output]].
+ */
+ def outputWithNames(names: Seq[String]): Seq[Attribute] = {
+// Save the output attributes to a variable to avoid duplicated
function calls.
+val outputAttributes = output
+assert(outputAttributes.length == names.length,
+ "The length of provided names doesn't match the length of output
attributes.")
+outputAttributes.zipWithIndex.map { case (element, index) =>
+ element.withName(names(index))
+}
+ }
+
--- End diff --
I was thinking...
```
object FileFormatWriter {
...
// workaround: a helper function...
def outputWithNames(outputAttributes: Seq[Attribute], names:
Seq[String]): Seq[Attribute] = {
assert(outputAttributes.length == names.length,
"The length of provided names doesn't match the length of output
attributes.")
outputAttributes.zipWithIndex.map { case (element, index) =>
element.withName(names(index))
}
}
```
Then, in each callsite, just say `FileFormatWriter.
outputWithNames(logicalPlan.output, names)`?
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user gengliangwang commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214646343
--- Diff:
sql/catalyst/src/main/scala/org/apache/spark/sql/catalyst/plans/QueryPlan.scala
---
@@ -38,6 +38,20 @@ abstract class QueryPlan[PlanType <:
QueryPlan[PlanType]] extends TreeNode[PlanT
*/
def outputSet: AttributeSet = AttributeSet(output)
+ /**
+ * Returns output attributes with provided names.
+ * The length of provided names should be the same of the length of
[[output]].
+ */
+ def outputWithNames(names: Seq[String]): Seq[Attribute] = {
+// Save the output attributes to a variable to avoid duplicated
function calls.
+val outputAttributes = output
+assert(outputAttributes.length == names.length,
+ "The length of provided names doesn't match the length of output
attributes.")
+outputAttributes.zipWithIndex.map { case (element, index) =>
+ element.withName(names(index))
+}
+ }
+
--- End diff --
It seems overkill to add a function here. But in `FileFormatWriter` we
can't not access `LogicalPlan` to get the attributes.
Another way is to put this method in a Util.
Do you have a good suggestion?
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user cloud-fan commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214645372
--- Diff:
sql/core/src/main/scala/org/apache/spark/sql/execution/datasources/DataSource.scala
---
@@ -495,7 +496,9 @@ case class DataSource(
s"Unable to resolve $name given
[${data.output.map(_.name).mkString(", ")}]")
}
}
-val resolved = cmd.copy(partitionColumns = resolvedPartCols,
outputColumns = outputColumns)
+val resolved = cmd.copy(
+ partitionColumns = resolvedPartCols,
+ outputColumnNames = outputColumns.map(_.name))
--- End diff --
why can't we use `outputColumnNames` directly here?
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user cloud-fan commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214644907
--- Diff:
sql/catalyst/src/main/scala/org/apache/spark/sql/catalyst/plans/QueryPlan.scala
---
@@ -38,6 +38,20 @@ abstract class QueryPlan[PlanType <:
QueryPlan[PlanType]] extends TreeNode[PlanT
*/
def outputSet: AttributeSet = AttributeSet(output)
+ /**
+ * Returns output attributes with provided names.
+ * The length of provided names should be the same of the length of
[[output]].
+ */
+ def outputWithNames(names: Seq[String]): Seq[Attribute] = {
+// Save the output attributes to a variable to avoid duplicated
function calls.
+val outputAttributes = output
+assert(outputAttributes.length == names.length,
+ "The length of provided names doesn't match the length of output
attributes.")
+outputAttributes.zipWithIndex.map { case (element, index) =>
+ element.withName(names(index))
+}
+ }
+
--- End diff --
or make it a util function
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user cloud-fan commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214644583
--- Diff:
sql/catalyst/src/main/scala/org/apache/spark/sql/catalyst/plans/QueryPlan.scala
---
@@ -38,6 +38,20 @@ abstract class QueryPlan[PlanType <:
QueryPlan[PlanType]] extends TreeNode[PlanT
*/
def outputSet: AttributeSet = AttributeSet(output)
+ /**
+ * Returns output attributes with provided names.
+ * The length of provided names should be the same of the length of
[[output]].
+ */
+ def outputWithNames(names: Seq[String]): Seq[Attribute] = {
+// Save the output attributes to a variable to avoid duplicated
function calls.
+val outputAttributes = output
+assert(outputAttributes.length == names.length,
+ "The length of provided names doesn't match the length of output
attributes.")
+outputAttributes.zipWithIndex.map { case (element, index) =>
+ element.withName(names(index))
+}
+ }
+
--- End diff --
+1
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user maropu commented on a diff in the pull request:
https://github.com/apache/spark/pull/22320#discussion_r214630233
--- Diff:
sql/catalyst/src/main/scala/org/apache/spark/sql/catalyst/plans/QueryPlan.scala
---
@@ -38,6 +38,20 @@ abstract class QueryPlan[PlanType <:
QueryPlan[PlanType]] extends TreeNode[PlanT
*/
def outputSet: AttributeSet = AttributeSet(output)
+ /**
+ * Returns output attributes with provided names.
+ * The length of provided names should be the same of the length of
[[output]].
+ */
+ def outputWithNames(names: Seq[String]): Seq[Attribute] = {
+// Save the output attributes to a variable to avoid duplicated
function calls.
+val outputAttributes = output
+assert(outputAttributes.length == names.length,
+ "The length of provided names doesn't match the length of output
attributes.")
+outputAttributes.zipWithIndex.map { case (element, index) =>
+ element.withName(names(index))
+}
+ }
+
--- End diff --
If #22311 merged, we don't need this function anymore? If so, IMHO it'd be
better to fix this issue in the `FileFormatWriter` side as a workaround?
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
Github user maropu commented on a diff in the pull request: https://github.com/apache/spark/pull/22320#discussion_r214609005 --- Diff: sql/core/src/main/scala/org/apache/spark/sql/execution/datasources/DataSource.scala --- @@ -460,9 +460,9 @@ case class DataSource( * @param mode The save mode for this writing. * @param data The input query plan that produces the data to be written. Note that this plan * is analyzed and optimized. - * @param outputColumns The original output columns of the input query plan. The optimizer may not - * preserve the output column's names' case, so we need this parameter - * instead of `data.output`. + * @param outputColumnNames The original output column names of the input query plan. The + * optimizer may not preserve the output column's names' case, so we need + * this parameter instead of `data.output`. --- End diff -- nit: ``` * @param outputColumnNames The original output column names of the input query plan. The * optimizer may not preserve the output column's names' case, so we need * this parameter instead of `data.output`. ``` --- - To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org For additional commands, e-mail: reviews-h...@spark.apache.org
[GitHub] spark pull request #22320: [SPARK-25313][SQL]Fix regression in FileFormatWri...
GitHub user gengliangwang opened a pull request:
https://github.com/apache/spark/pull/22320
[SPARK-25313][SQL]Fix regression in FileFormatWriter output names
## What changes were proposed in this pull request?
Let's see the follow example:
```
val location = "/tmp/t"
val df = spark.range(10).toDF("id")
df.write.format("parquet").saveAsTable("tbl")
spark.sql("CREATE VIEW view1 AS SELECT id FROM tbl")
spark.sql(s"CREATE TABLE tbl2(ID long) USING parquet location
$location")
spark.sql("INSERT OVERWRITE TABLE tbl2 SELECT ID FROM view1")
println(spark.read.parquet(location).schema)
spark.table("tbl2").show()
```
The output column name in schema will be `id` instead of `ID`, thus the
last query shows nothing from `tbl2`.
By enabling the debug message we can see that the output naming is changed
from `ID` to `id`, and then the `outputColumns` in
`InsertIntoHadoopFsRelationCommand` is changed in `RemoveRedundantAliases`.
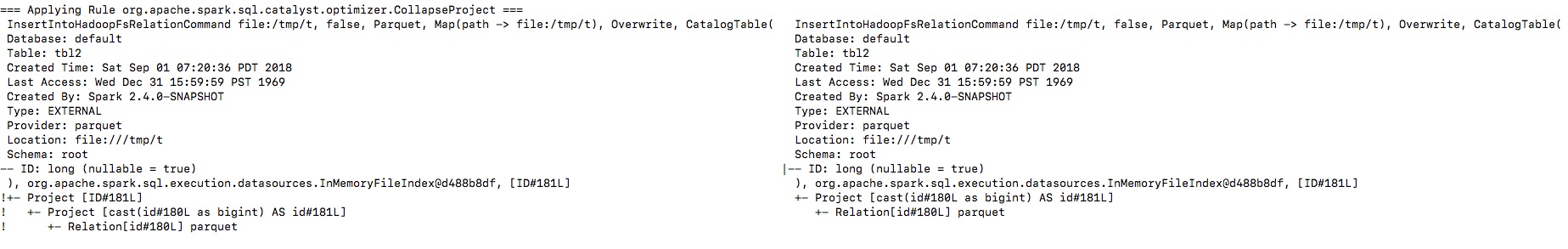
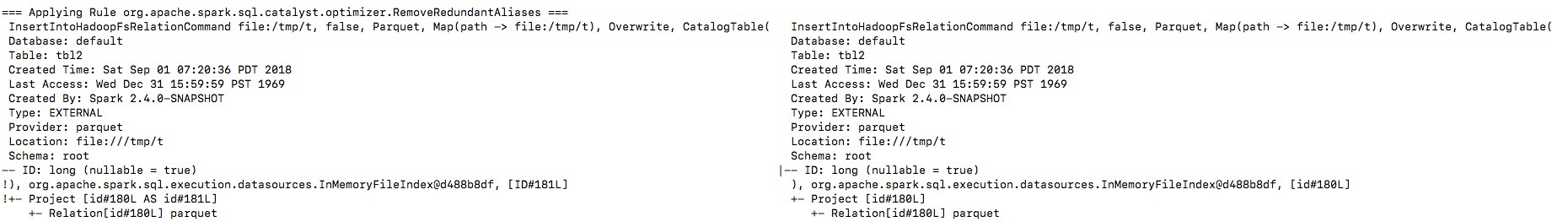
**To guarantee correctness**, we should change the output columns from
`Seq[Attribute]` to `Seq[String]` to avoid its names being replaced by
optimizer.
I will fix project elimination related rules in
https://github.com/apache/spark/pull/22311 after this one.
## How was this patch tested?
Unit test.
You can merge this pull request into a Git repository by running:
$ git pull https://github.com/gengliangwang/spark fixOutputSchema
Alternatively you can review and apply these changes as the patch at:
https://github.com/apache/spark/pull/22320.patch
To close this pull request, make a commit to your master/trunk branch
with (at least) the following in the commit message:
This closes #22320
commit bbd572c1fe542c6b2fd642212f927ba384c882e4
Author: Gengliang Wang
Date: 2018-08-31T16:07:00Z
Fix regression in FileFormatWriter output schema
---
-
To unsubscribe, e-mail: reviews-unsubscr...@spark.apache.org
For additional commands, e-mail: reviews-h...@spark.apache.org
